DNA Wheatfen
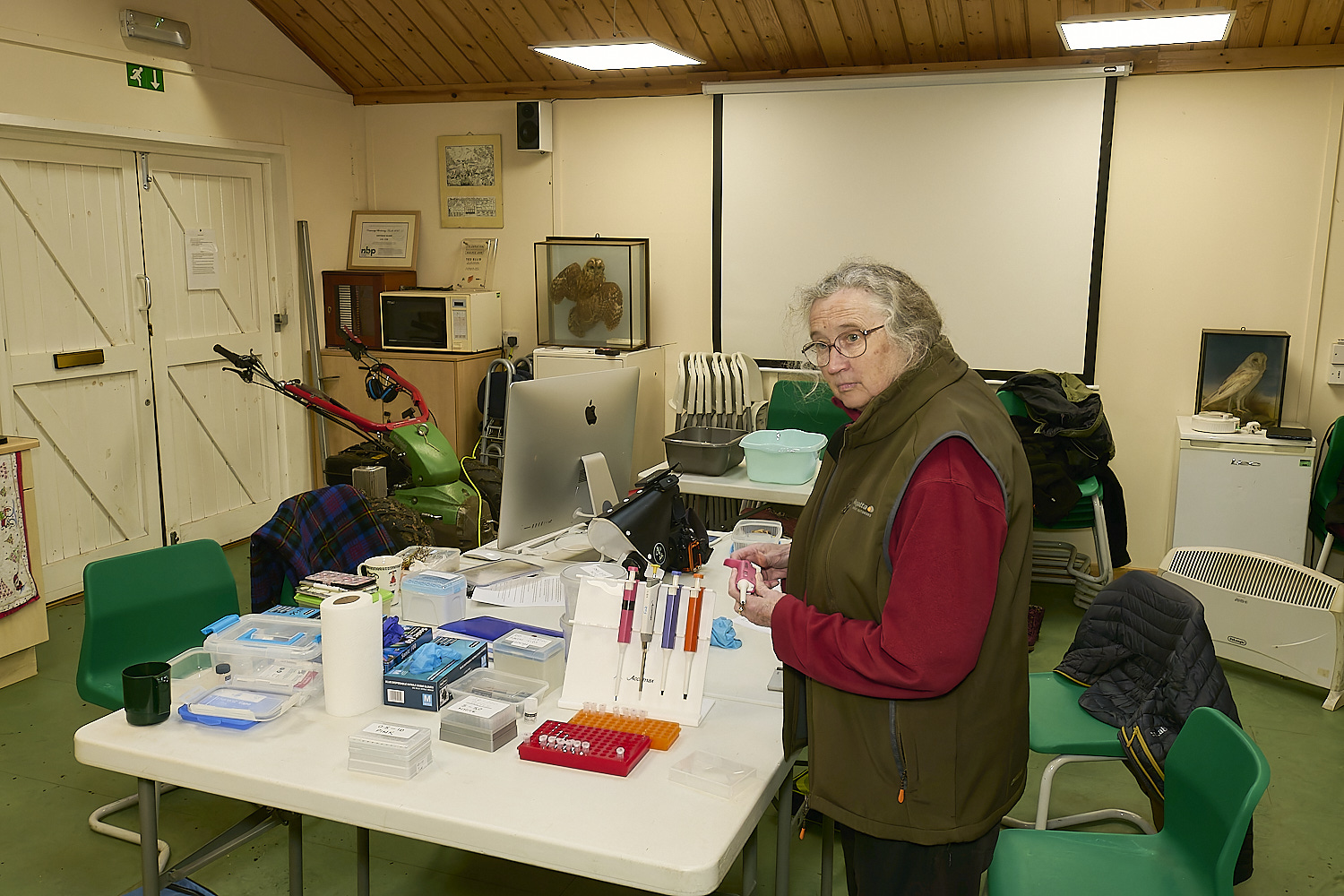
The Norfolk Fungus Study Group
DNA team
at work at
Wheatfen
under the watchful gaze of
Wheatfen's founder
Ted Ellis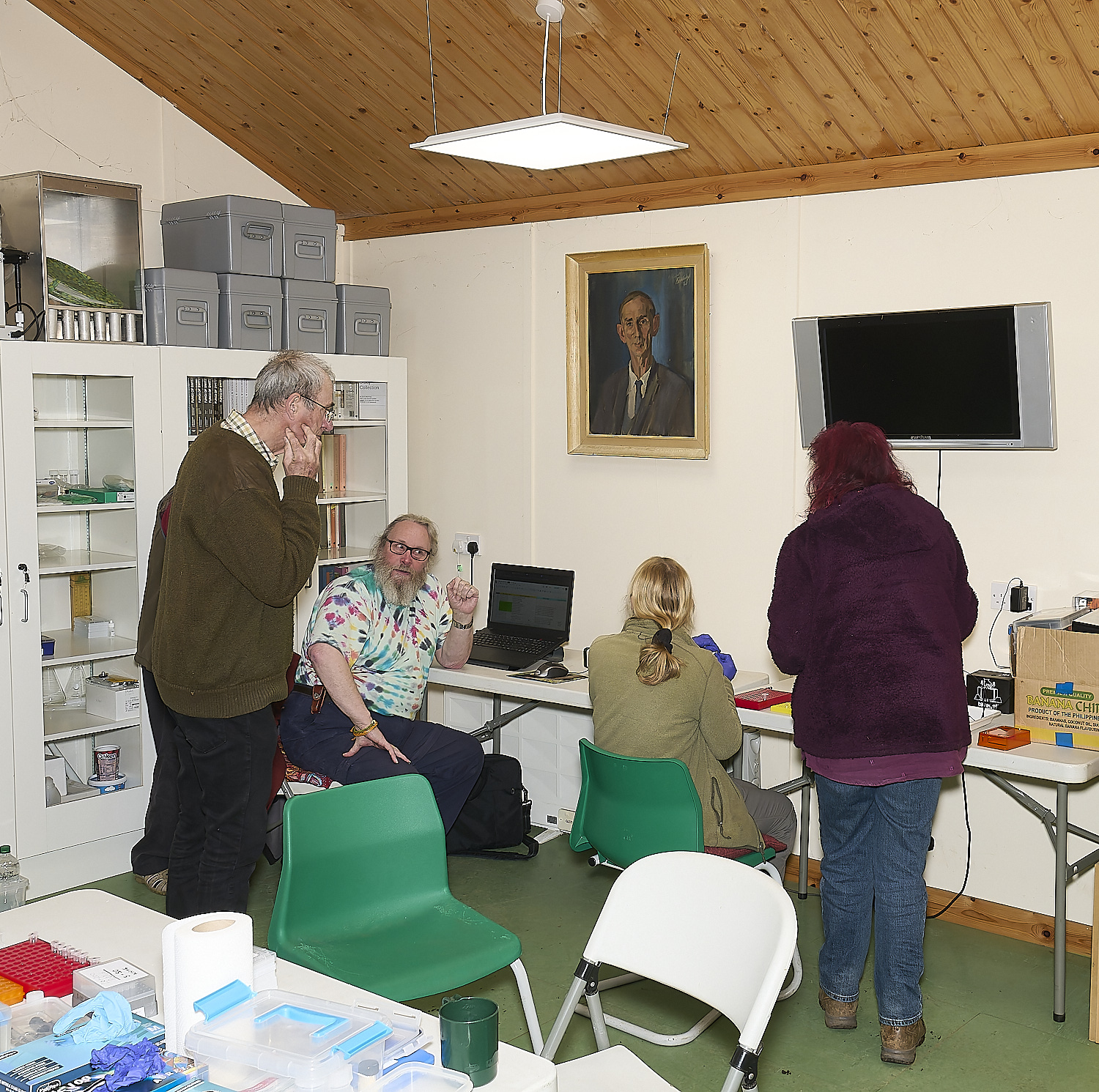
Working surfaces are cleaned and sterilised with 10% thin domestic bleach in tap water with a drop of washing up liquid.
This is vital to prevent contamination, particularly if there of risk of amplified DNA on the surfaces.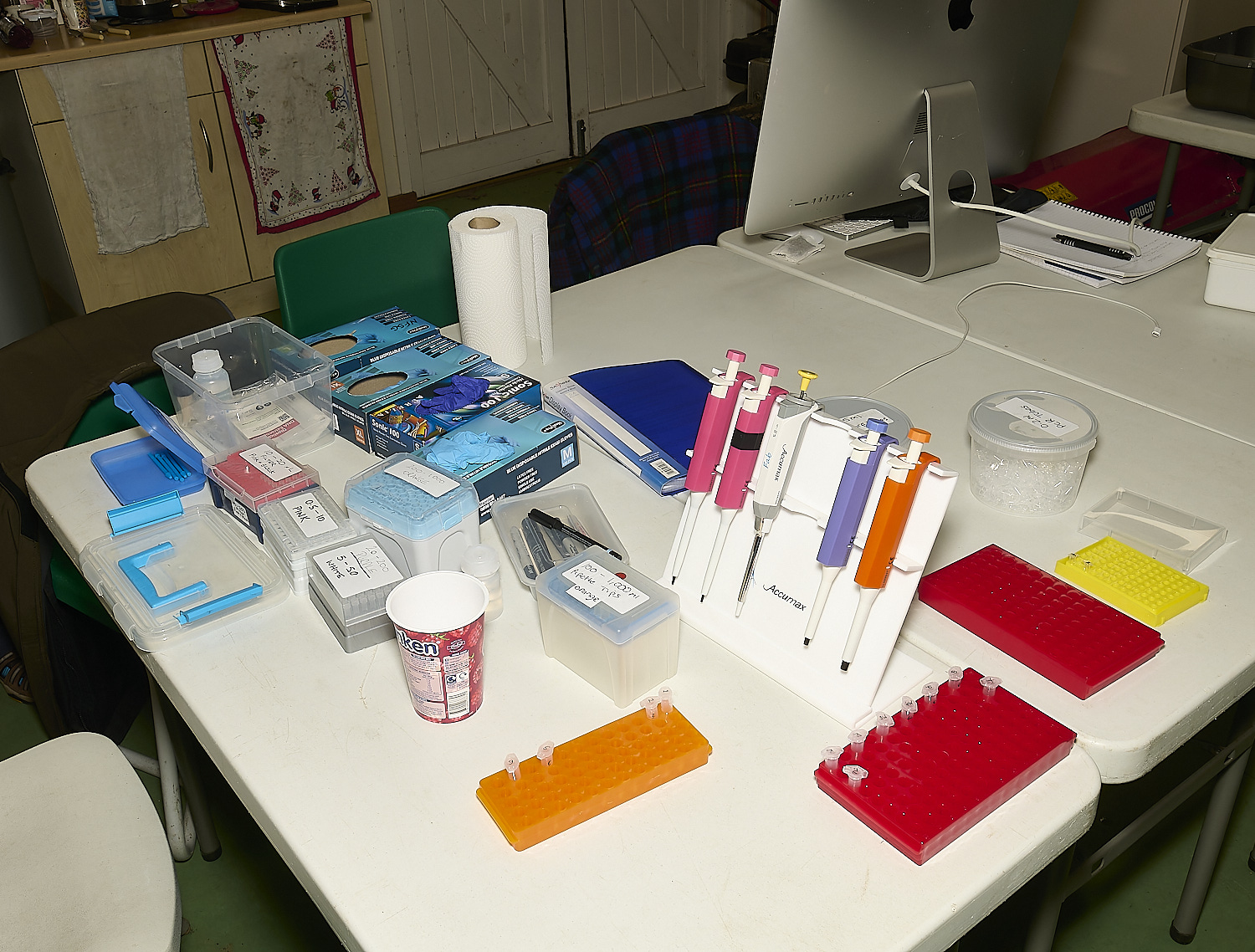
A variety of pipettes, epindorf tubes & gloves needed for the process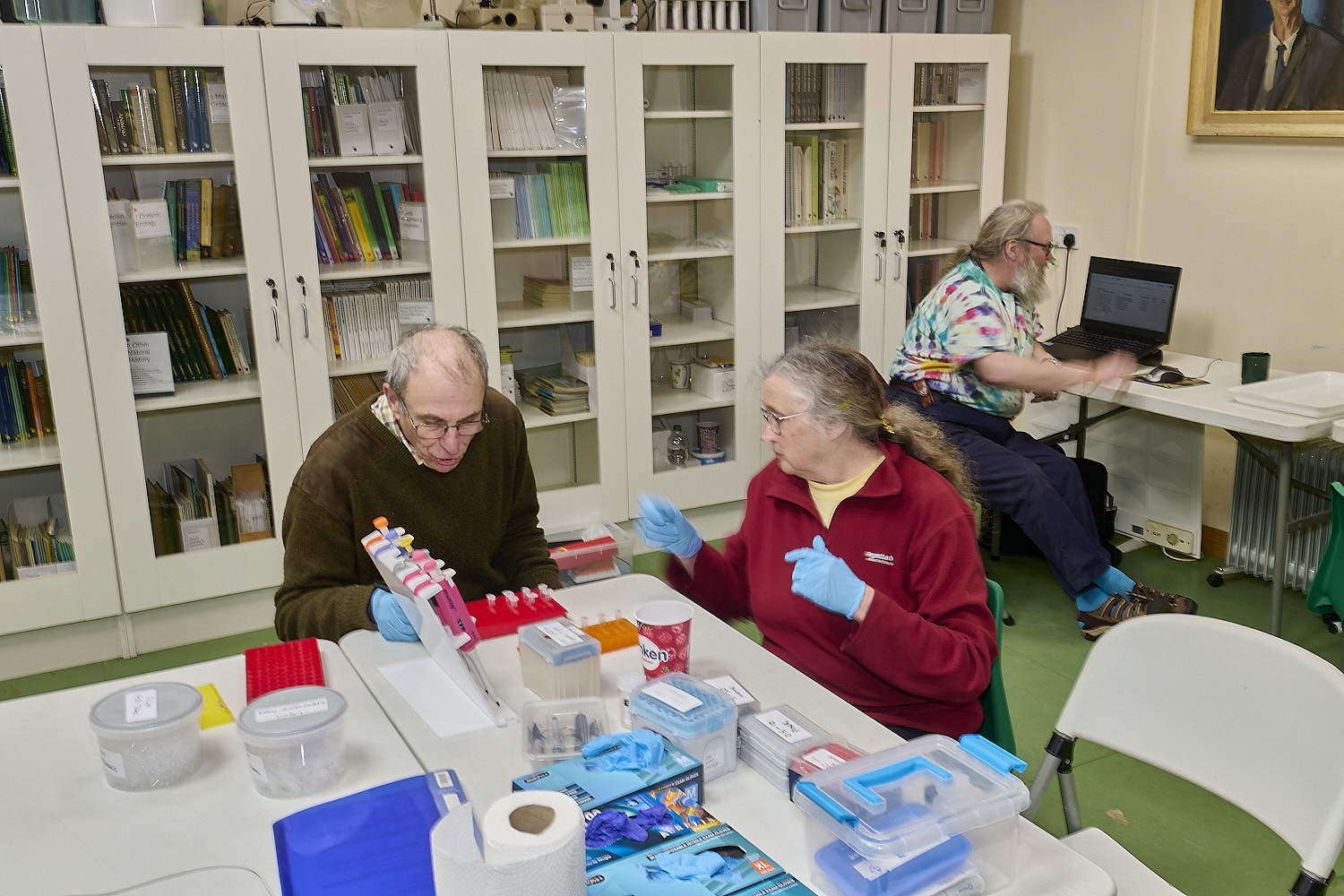
Preparing numbered 1.5ml epindorf tubes with 100µl of extraction buffer ready for the samples.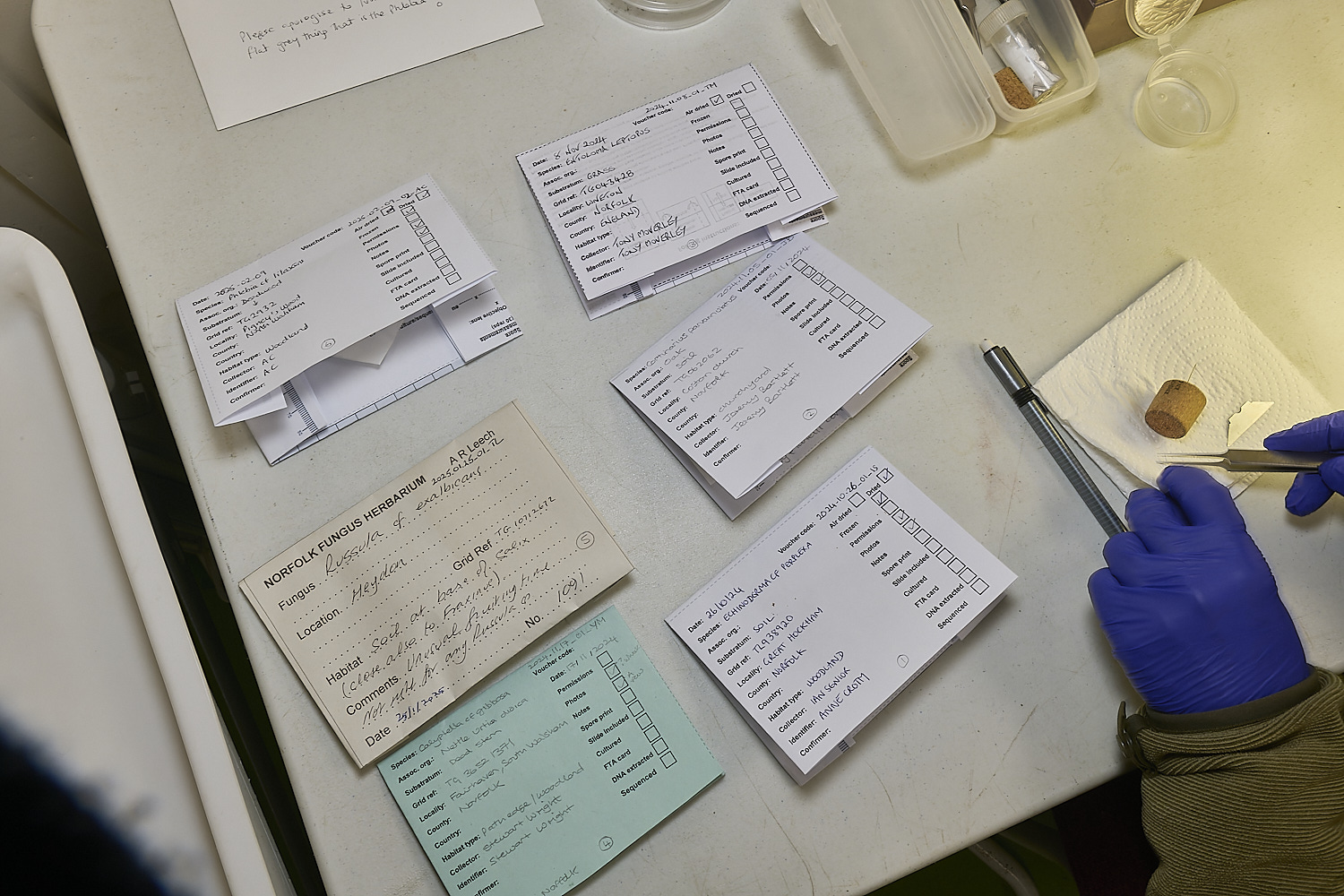
Fungal specimens submitted in labelled packets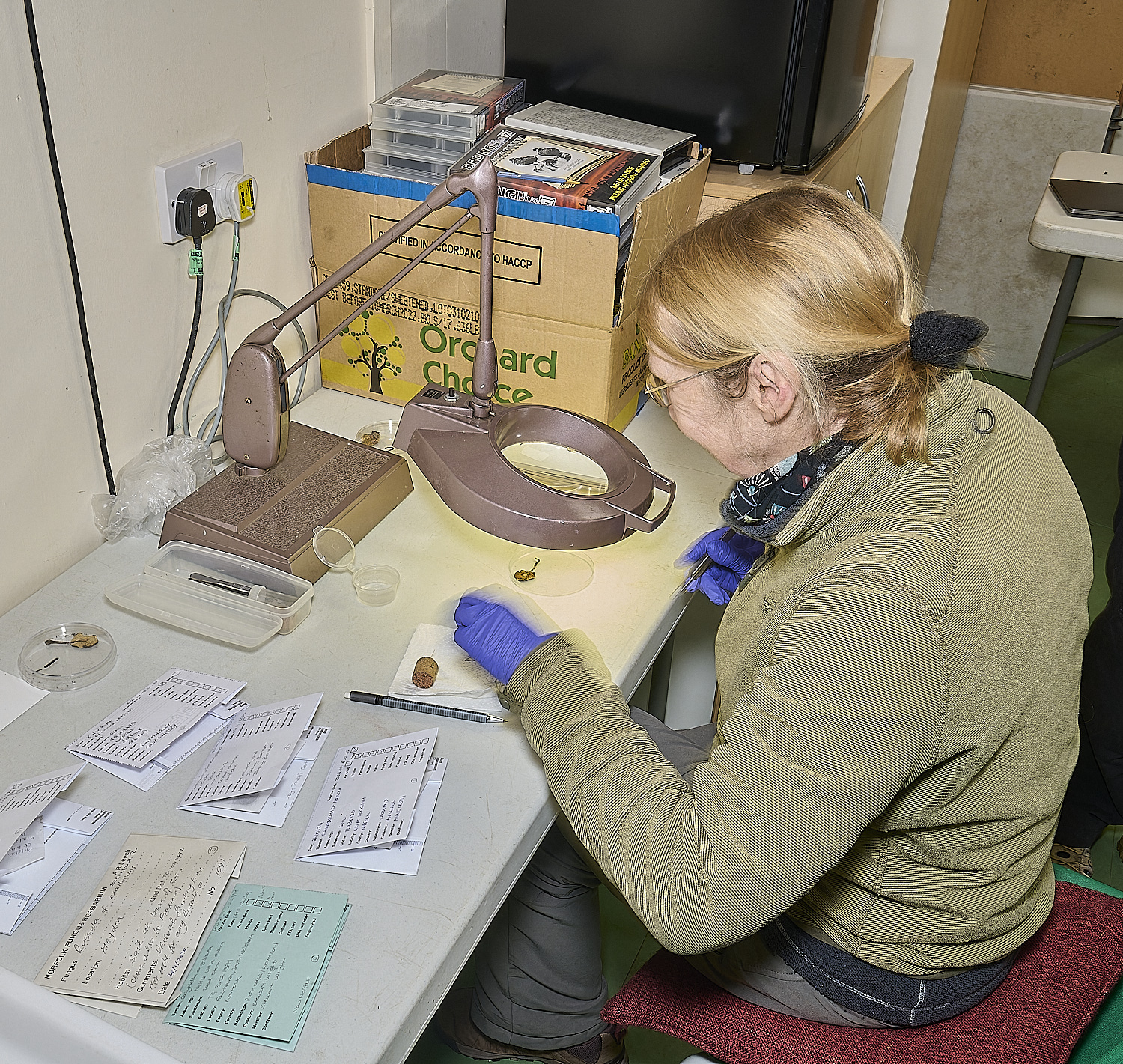
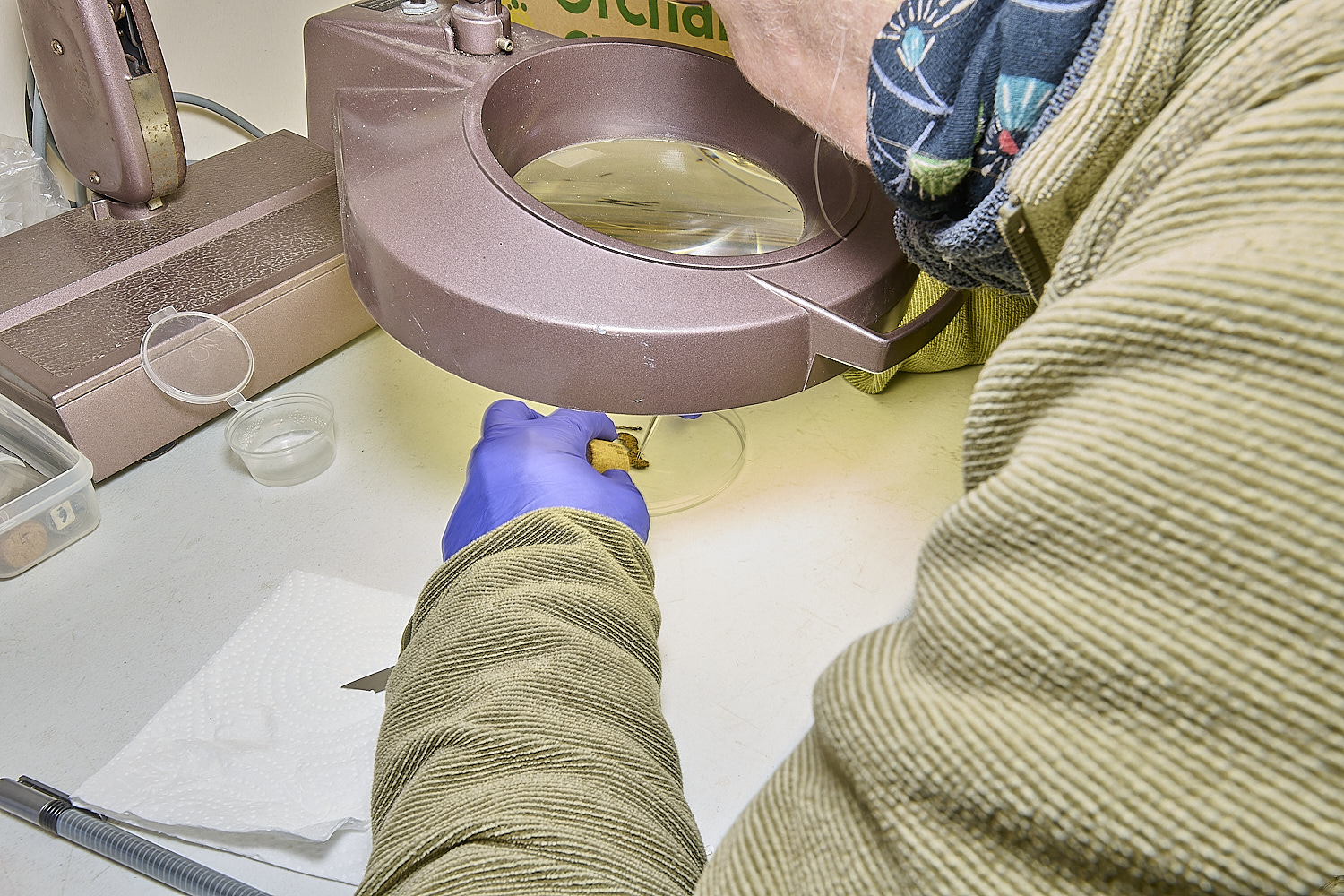
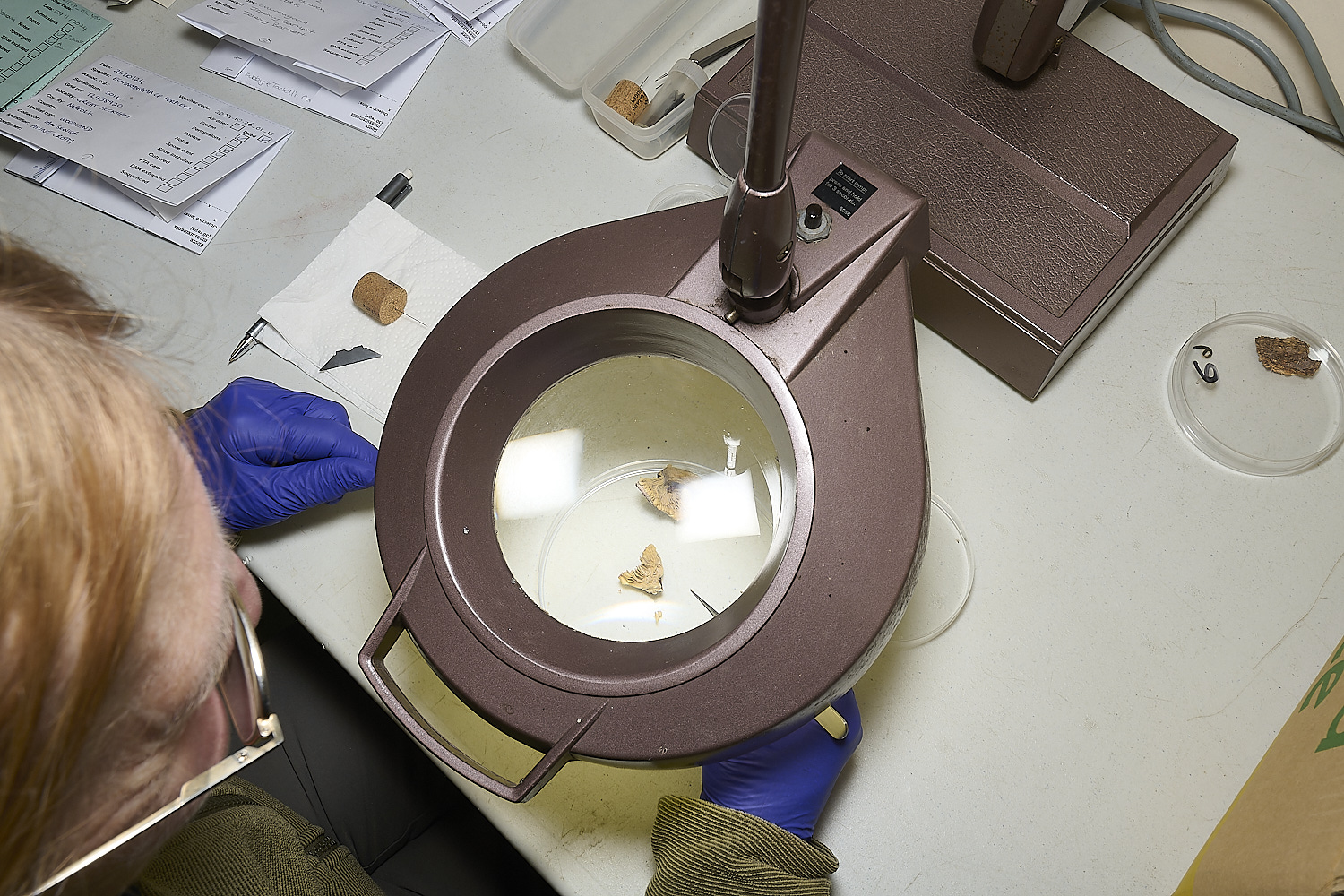


Using sterilised tweezers, razor blades and needles, and sterilising instruments in bleach between specimens,
small samples of fungal tissue are cut or torn off from the samples for testing.
Each sample is then placed in an appropriately numbered 1.5ml epindorf tube containing extraction buffer as mentioned above.
The samples are then ground with a sterile (bleached pestle) until the tissue is mostly pulverised.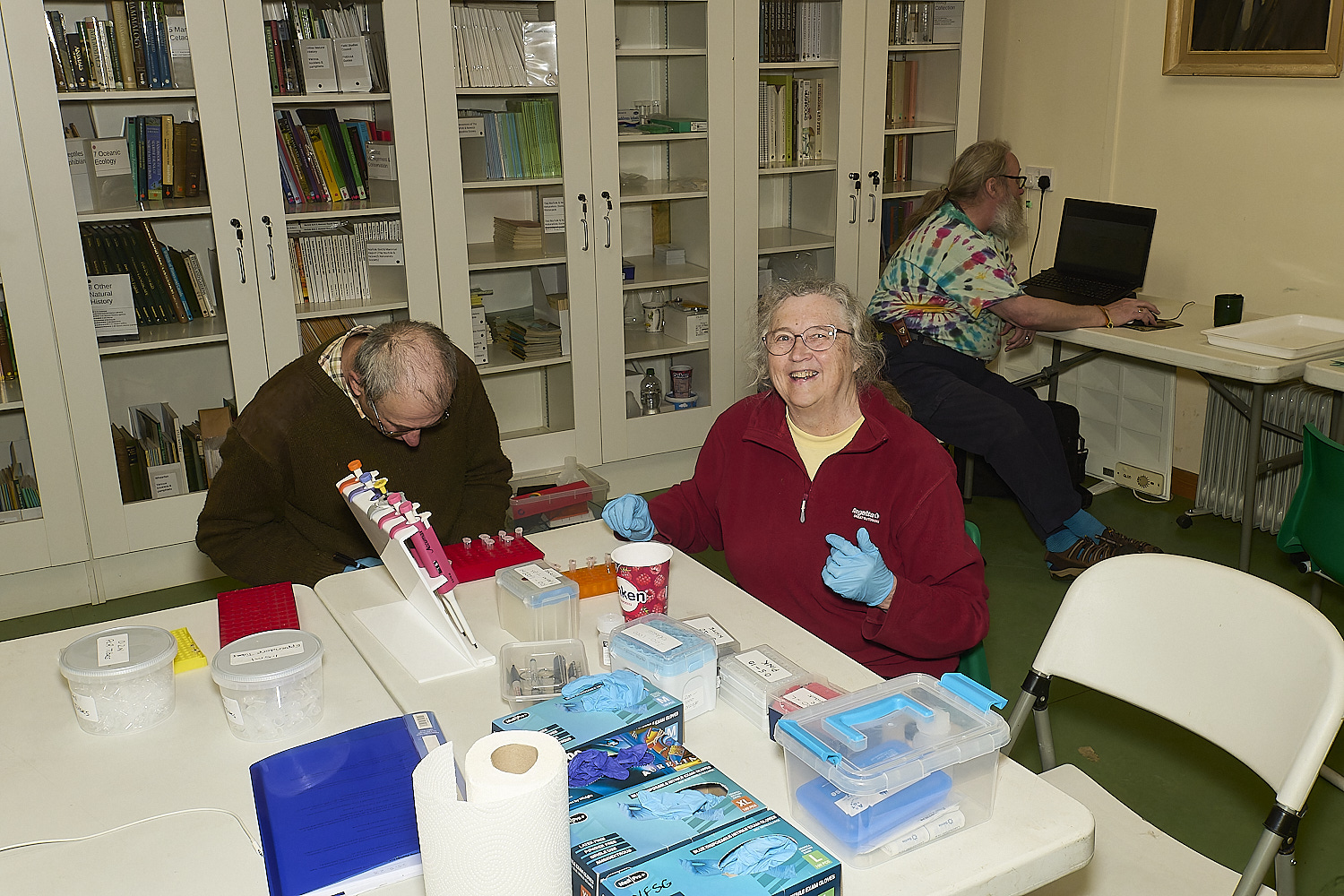
Once ground teach tube has a further 400µl of extraction buffer added to it.
A seperate set of similarly numbered tubes have 1000µl of wash Buffer added to them.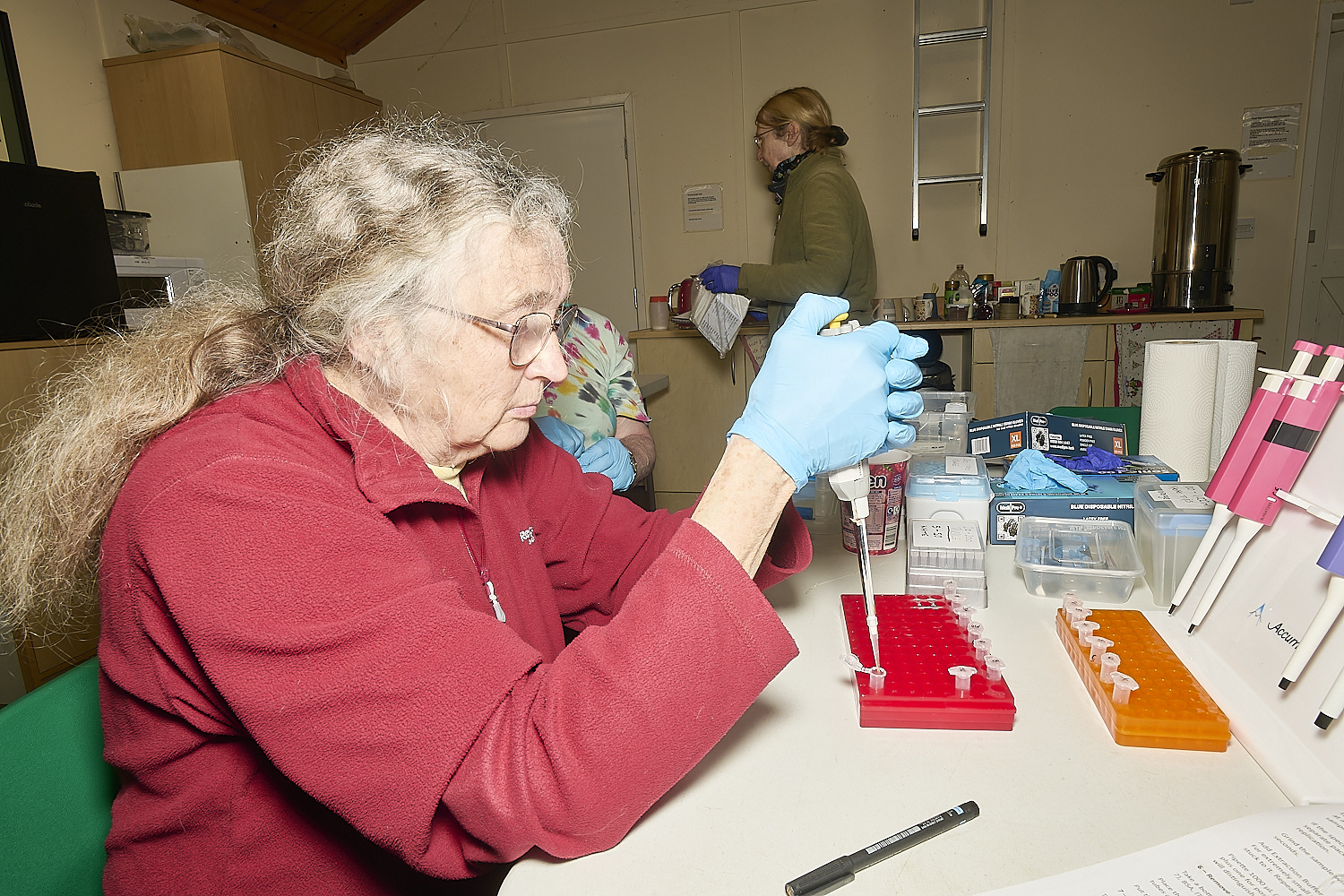
Preparation of the 5x HOT FIREPol Master MIx
152µl of PCR Grade Water
40µl 5x HOT FIREPol Mix
4µl of prepared ITS1F primer at working concentration
4µl of prepared ITS4 primer at working concentration
Numbered tubes are then prepared, each containing 20µl of this solution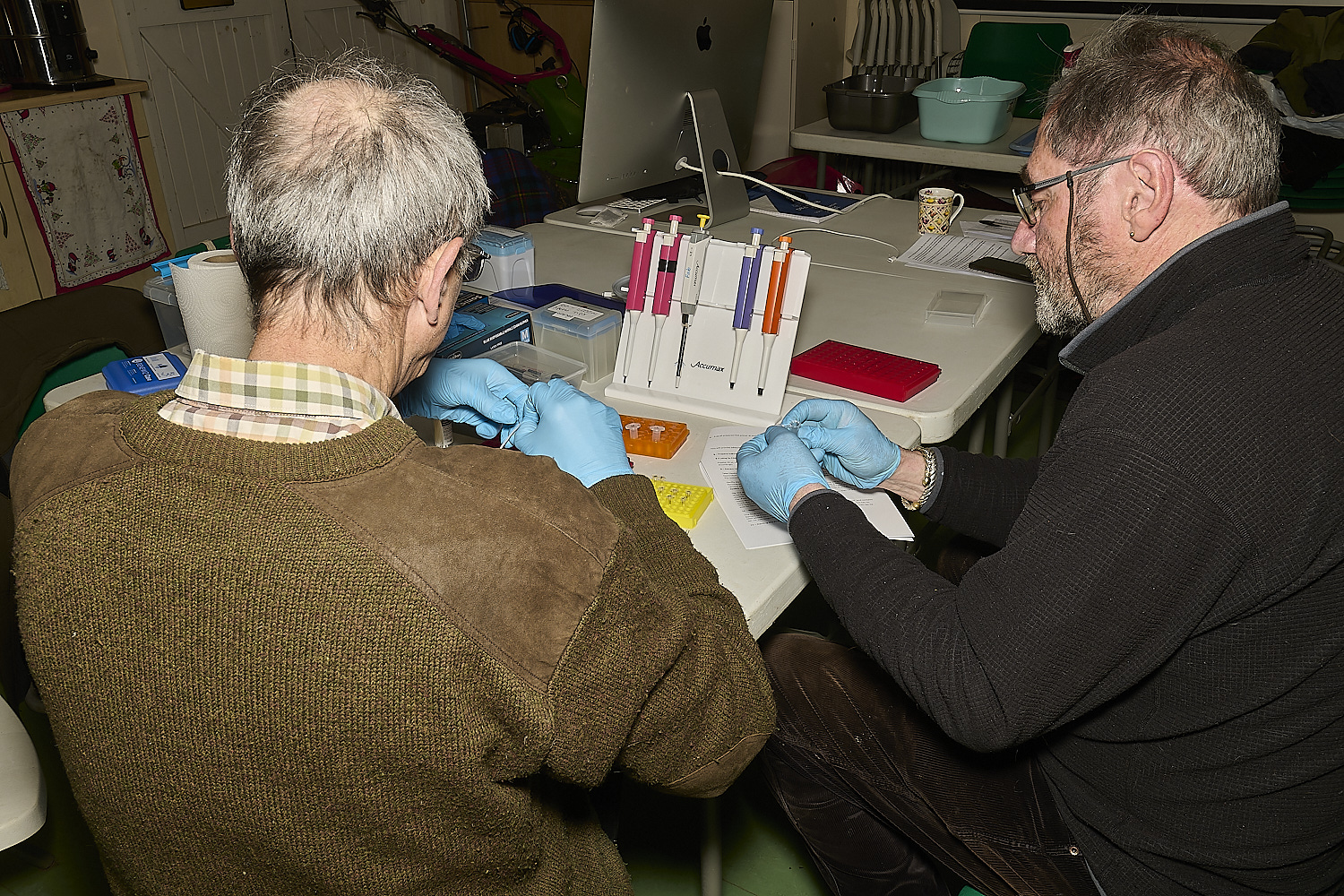
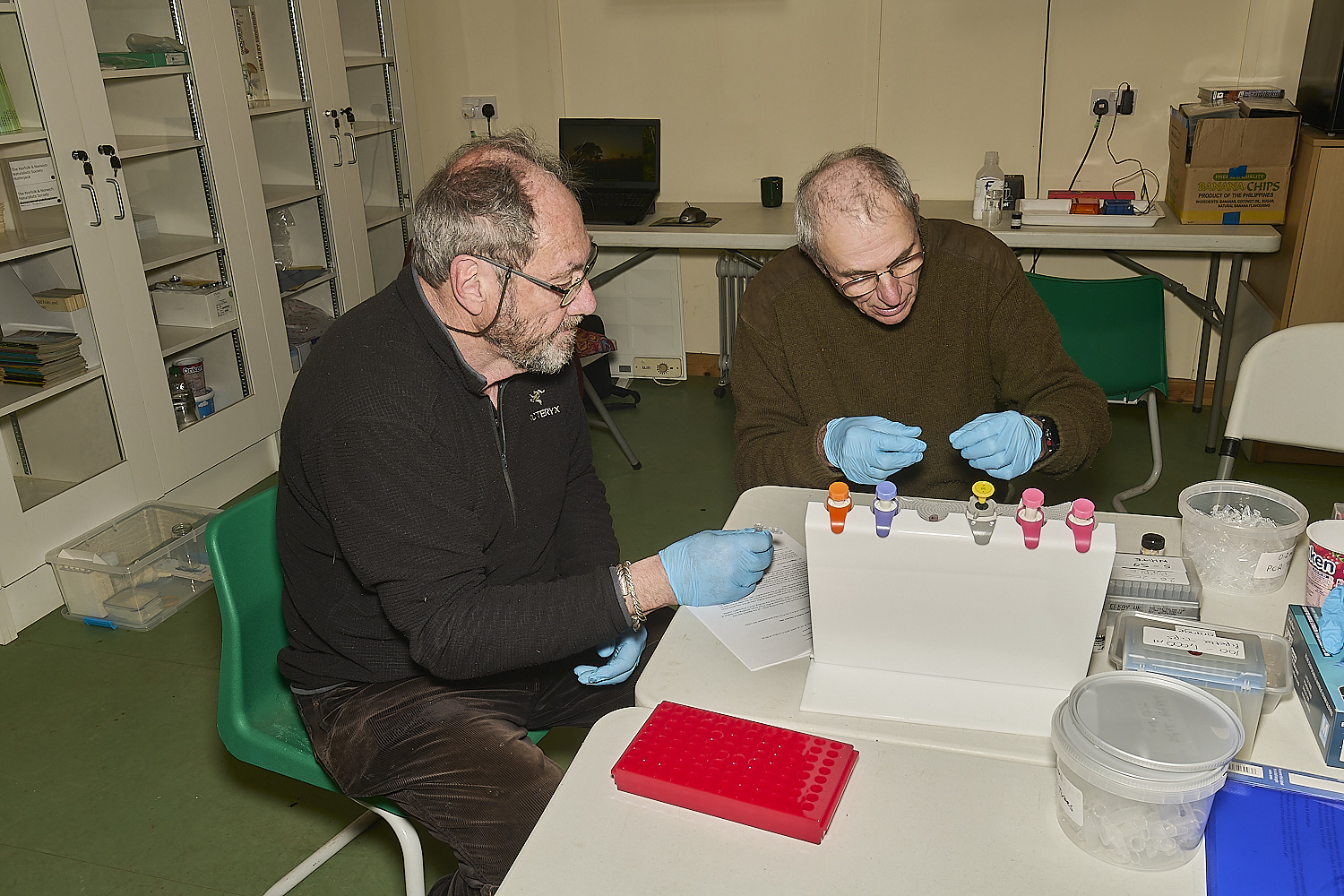

This is the moment when it all comes together and DNA is extracted.
1 A dipstick is dipped into the epindorf tube containing the sample with the extraction buffer three times.
2 The dipstick is then dipped into the tube containing the wash buffer three times
3 The dipstick is then dipped into the PCR mix tube three times to release the DNA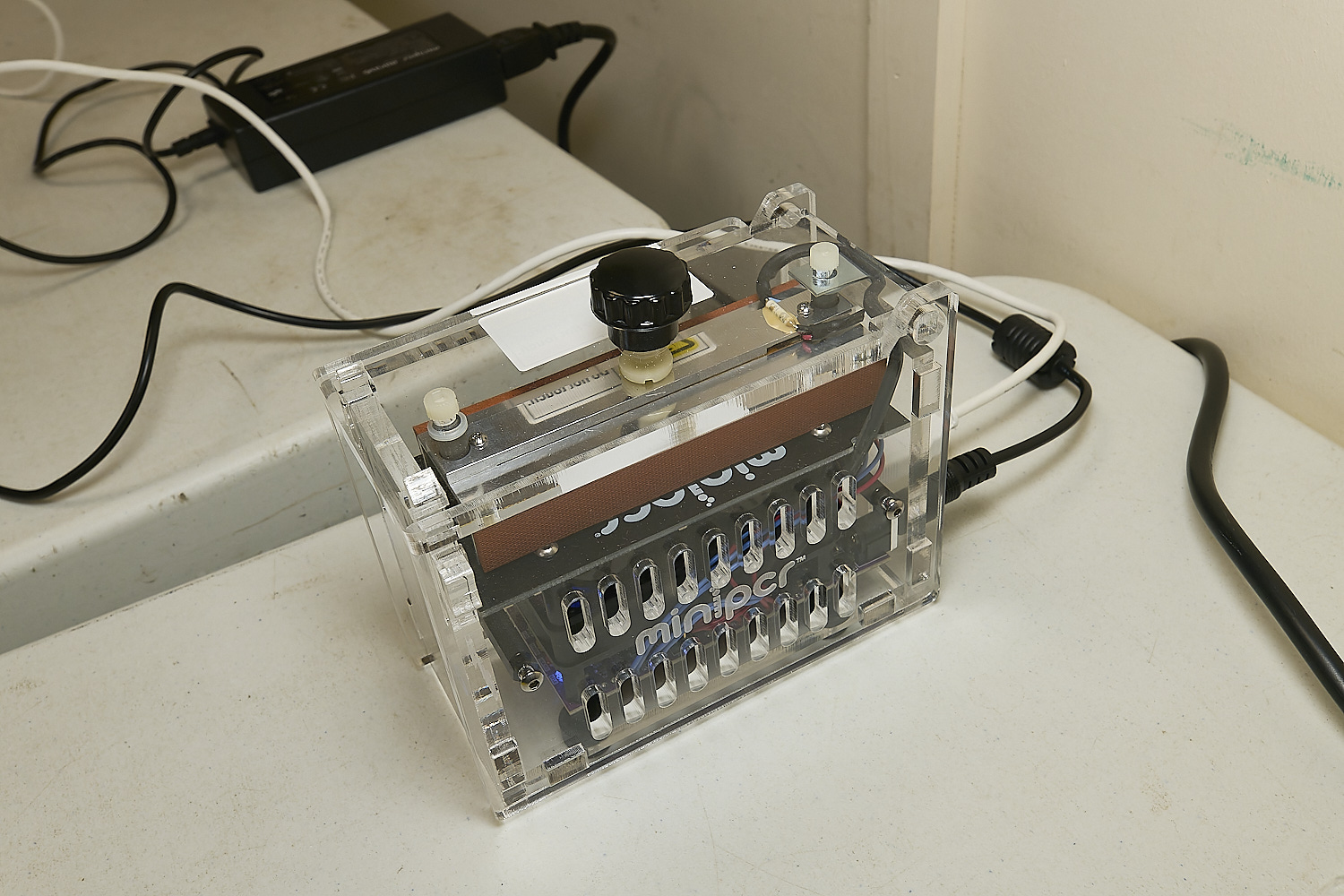
The PCR machine where the prepared samples complete with primers and FIREPol Mastermix are put through a thermal cycle
to amplify the DNA present.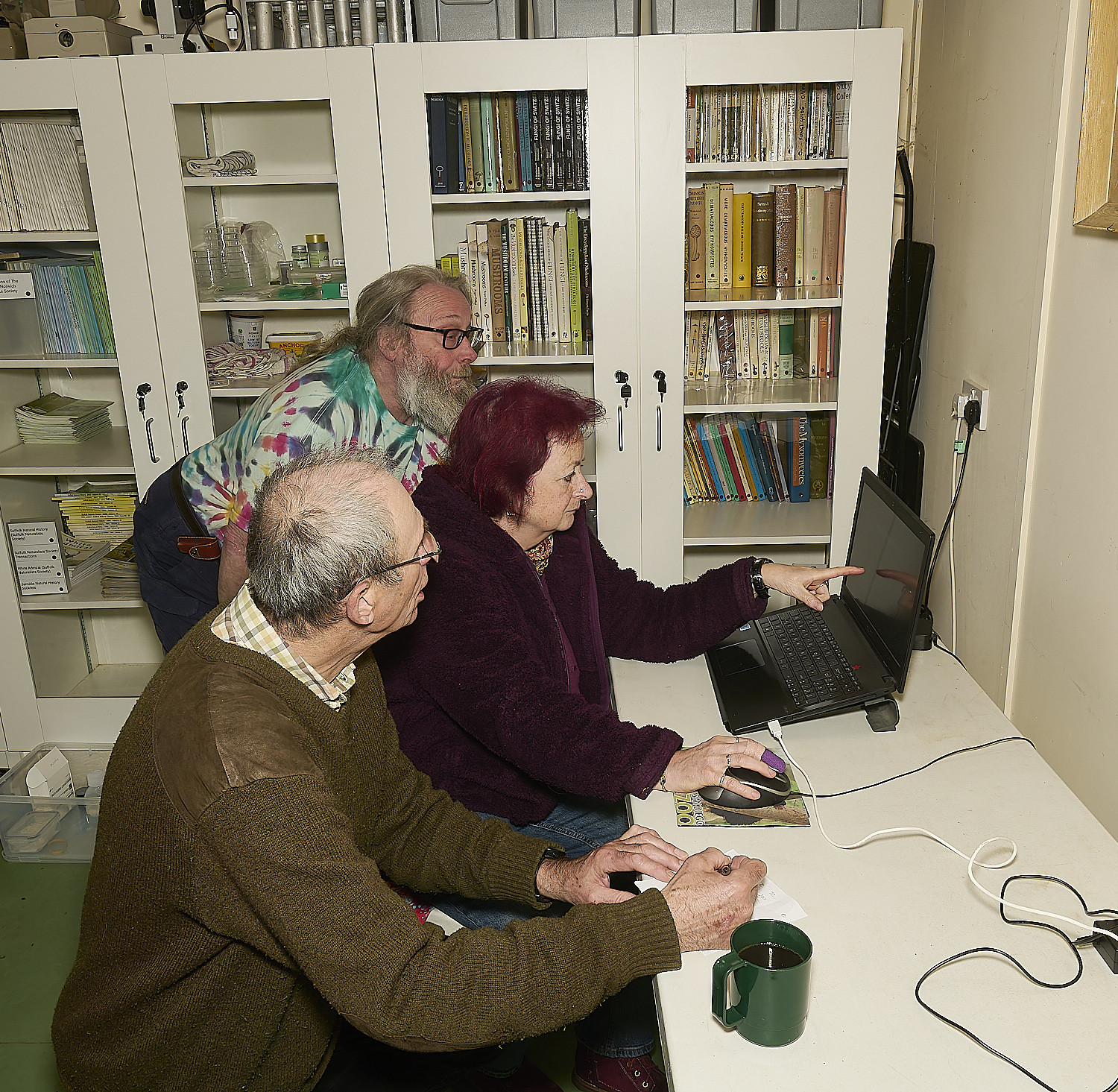
Lab book recording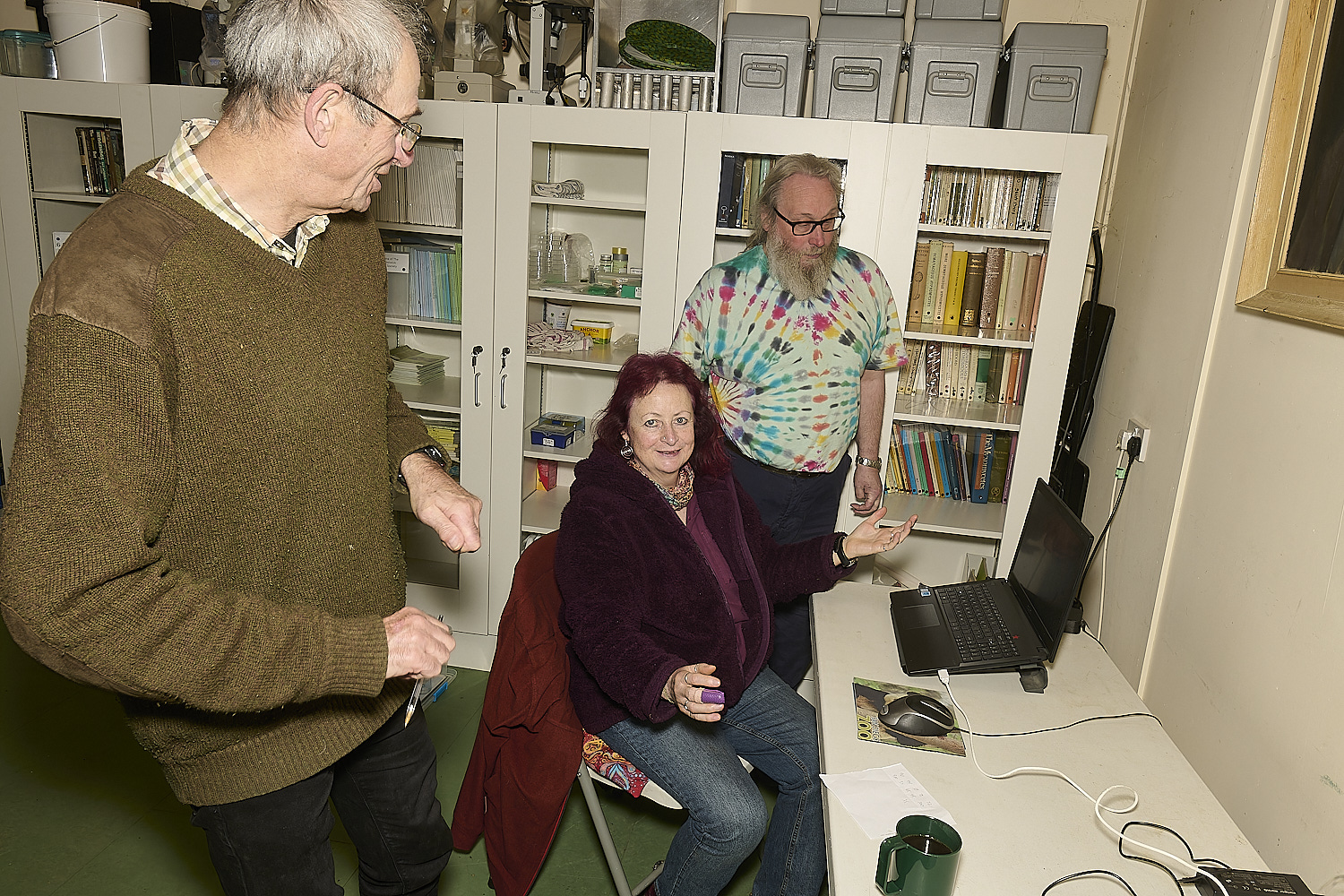
We've done it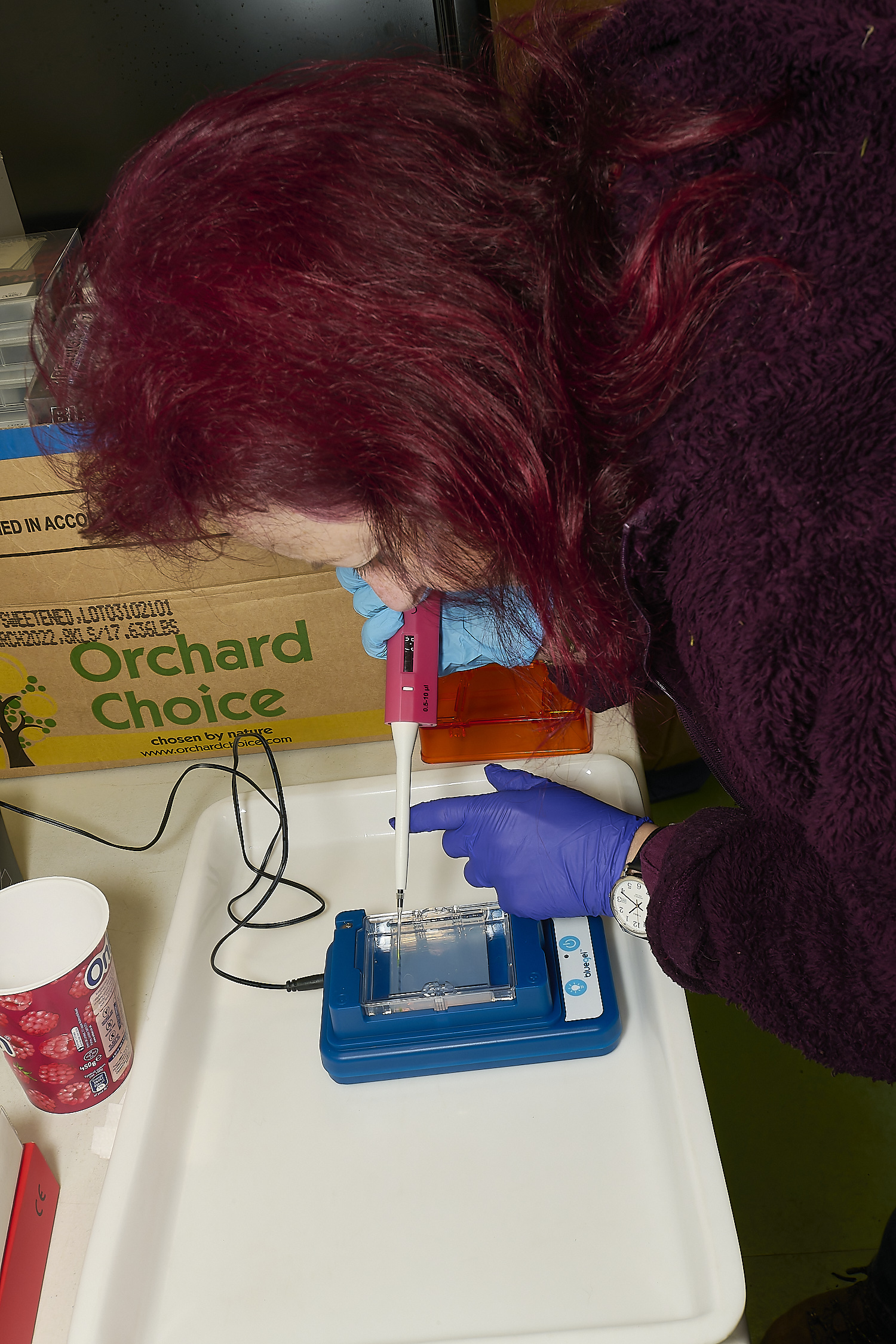
Preparation for GEL electrophoresis
